SPRm200 Surface Plasmon
Resonance Microscopy is based on surface plasmon resonance (SPR) technology,
which is the first biosensor integrating
optical microscopy with Surface Plasmon Resonance technology around the
world. SPRm200 can obtain cell in
situ bright imaging、 SPR imaging、 SPR kinetic curve of quantitative affinity
constant and binding dissociation constant and it opens a new frontier in the
label-free study of molecule interaction. SPRm200 designed to label-free detect
on cell membrane proteins and related molecule enables you not only not to
extract or separate membrane proteins but also monitor the cell structure and
measure the binding process of drug and target simultaneously.Meanwhile,
SPRm200 can also measure the interaction of crude membrane proteins with drug
and high resolution imaging can also be realized.Additionally, SPRm200 make it
possible to realize the study of single or multiple cell, as a result, it can
realize the heterogeneity of cell and drug binding and statistical
analysis. Due to its outstanding
sensitivity and stability, SPRm200 can measure the binding behaviour in
nanoscale, which can study the interaction of bacteria or virus with antibacterial
drug and develop new nanoparticles delivered by drugs.

(1) Binding screening and analysis of micromolecule drug
and single cell
(2) Binding screening of micromolecule drug(<200Da) and multiple or living cells and
precision statistical distribution analysis of single molecule, study on
heterogeneity differences of cell
(3) Binding screening of antibody drug and multiple or single cells and precision statistical
distribution analysis of single molecule, study on heterogeneity differences of
cell
(4) Interactions of bacteria or virus with antibacterial
drug
Study
on layer in situ of other molecule cell or living cell
Main
function
Quantitative mapping of the interaction of living cell
molecular in real time
Kinetic & quantitative analysis and statistical
distribution of single or multiple cells
Bright field imaging、 SPR imaging combined with kinetic analysis in single
point
Biomarker Monitoring, Drug target development
Label-free micromolecule analysis(<200Da)
Electrochemical
analysis
Micromolecule、DNA、antibody、peptide、protein、virus、bacteria、cell
Technology features
Cell in situ: realize the in situ molecule
interaction of cell level, not need to purify, in situ analyze the interaction
of drug with molecule in cell level, especially provide solution to membrane
proteins receptor (glycoprotein 、GPCR) and so on;
High flux: SPR field of view
is 600um*600um, so it can study the interaction of multiple cell and molecule
simultaneously.As a result, it can realize the drug statistical analysis and
cell heterogeneity study.
High precision: SPRm200 can realize the
study on interaction of micromolecule(<200Da) with cells;
High resolution dynamic visualization:
SPRm200 can monitor the whole process
and imaging of the interaction of drug molecule and cell in a dynamic and
visualized way.In the meanwhile, it can obtain the quantitative SPR
analysis curve and obtain quantitative
Ka,Kd ,KD. The high solution(1 um) imaging can realize single cell’s solution imaging and
quantitative data.
3.Technology abilities
Base Station | Light source | 690nm |
Field of view | Bright Field:1200*900um SPR:600*450um |
Resolution | 1um |
Baseline noise | <0.6RU RMS(0.1 mDeg RMS) |
Fluid Handling | Sample volume | 1-1500ul |
Sample stage translation/rotation | 3mm*3mm/360 deg |
Solution tranferring method | Semi-auto/Auto |
Minimum injection time | <0.2s |
Molecular weight cutoff | 200 Da |
Study based on in situ cell molecule
interaction
Micromolecule drug
Among common drugs, micromolecule
drug represents 98% of the total. Micromolecule drug is usually a inhibitor
delivering signal and it can specifically block the necessary signal conducting
pathway in the process of tumor growth and proliferation and finally it can
realize the aim of treatment. Micromolecule drug provides powder to be absorbed
through concentration gradient.
1.Interaction of micromolecule with HEK
293 cell and GPR39 receptor
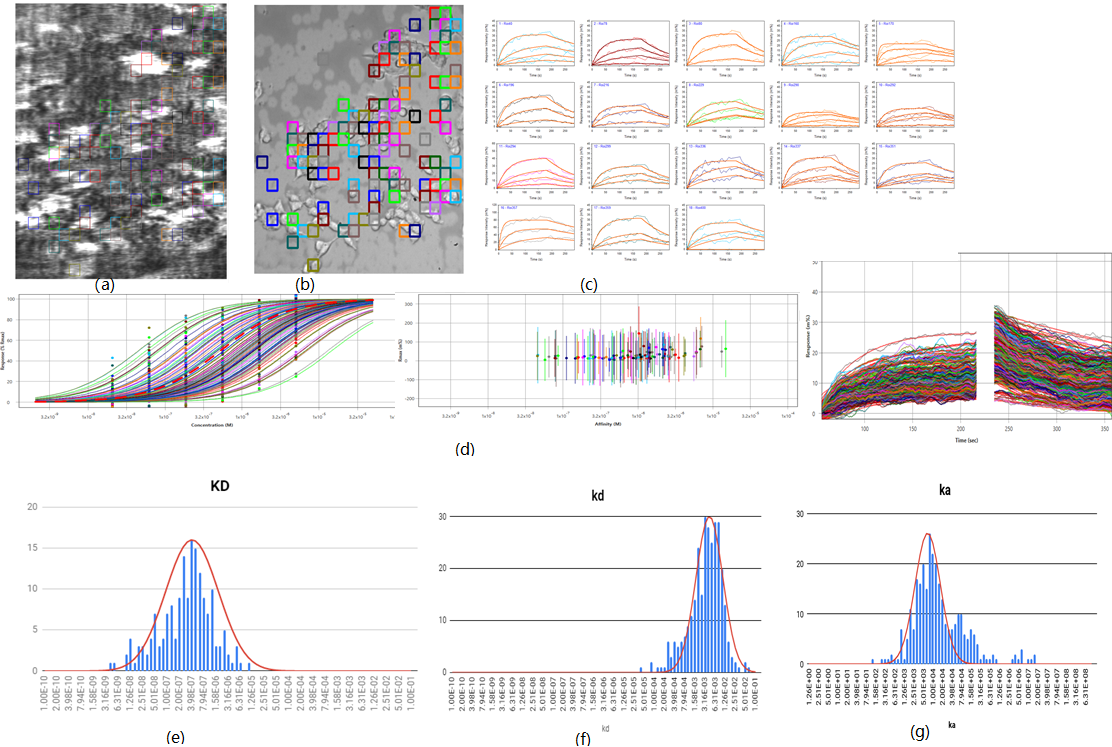
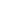
Fig.(a)SPR imaging,brightness and
darkness reflect the degree of interaction among different cell and district ;
Fig.(b), bright imaging, can obtain the observation and
district of different cell; Fig.(c)multiple interest
district, SPR signal curve in every district, Fig(e,f,g) multiple interest
district respectively,statistical distribution of affinity constant/dissociation constant/biding constant:KD:413nM(42nM standard deviation),kd = 4.27E-3
s-1 (0.720E-3 standard deviation),ka = 6.92E3 M-1 s-1 (0.721E3
standard deviation)
2.Study on micromolecule and interaction
of cell ASIC acid sensitive ionic pathway receptor
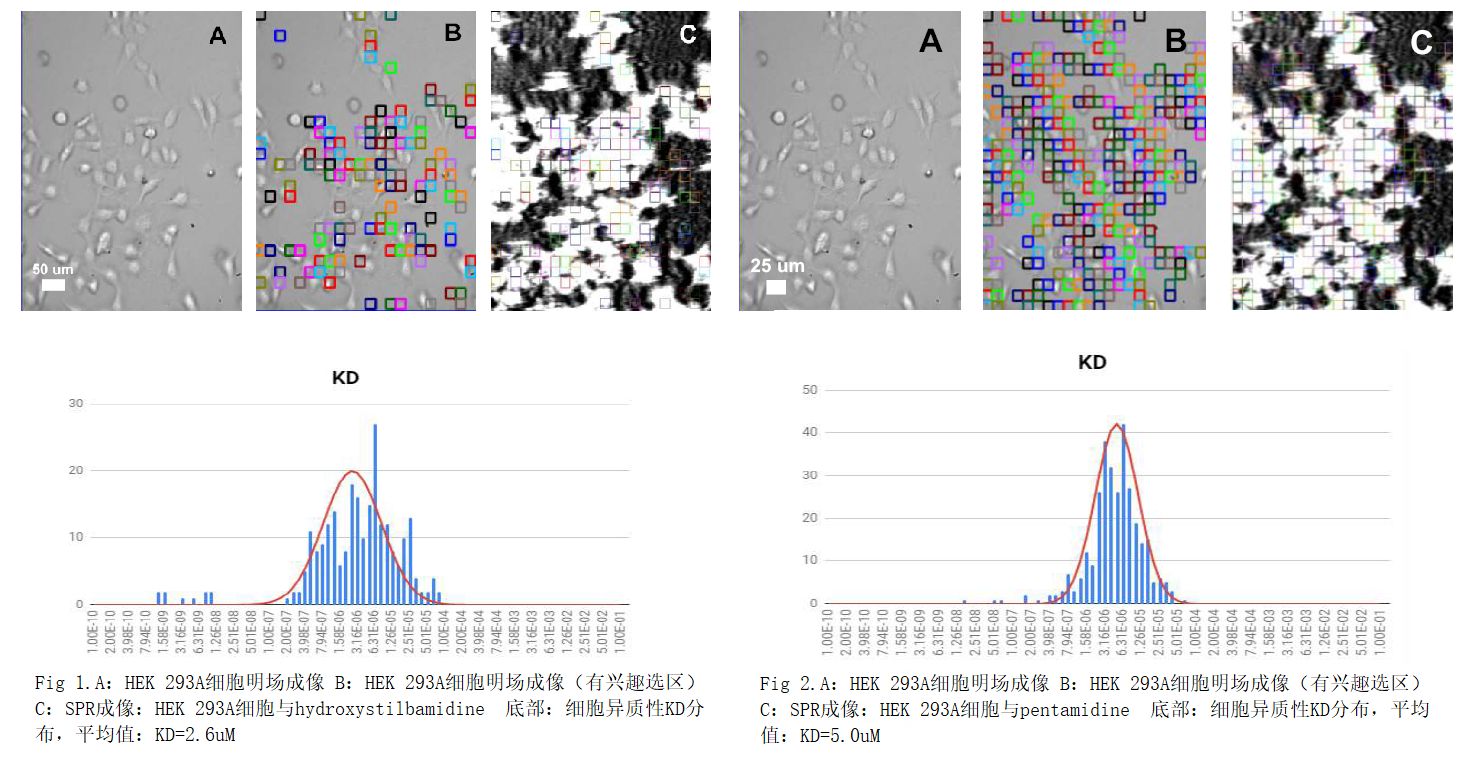
Antibody drug
Monoclonal antibody (mAb)therapy has became a ripe
method to treat cancer ,andautoimmune disease,
asthma and many other diseases.Monoclonal antibody (mAb)drug account for 50% of the whole
biopharmacy , more than 60% of which are membrane protein receptors. SPRm200
allows quantitative studies of monoclonal antibody (mAb) and membrane protein
binding at the single-cell level.Traditional SPR directly fixes the purified
protein on the chip surface, which has problems in membrane proteins and it is difficult
to extract from the cell environment and maintain their own structure.
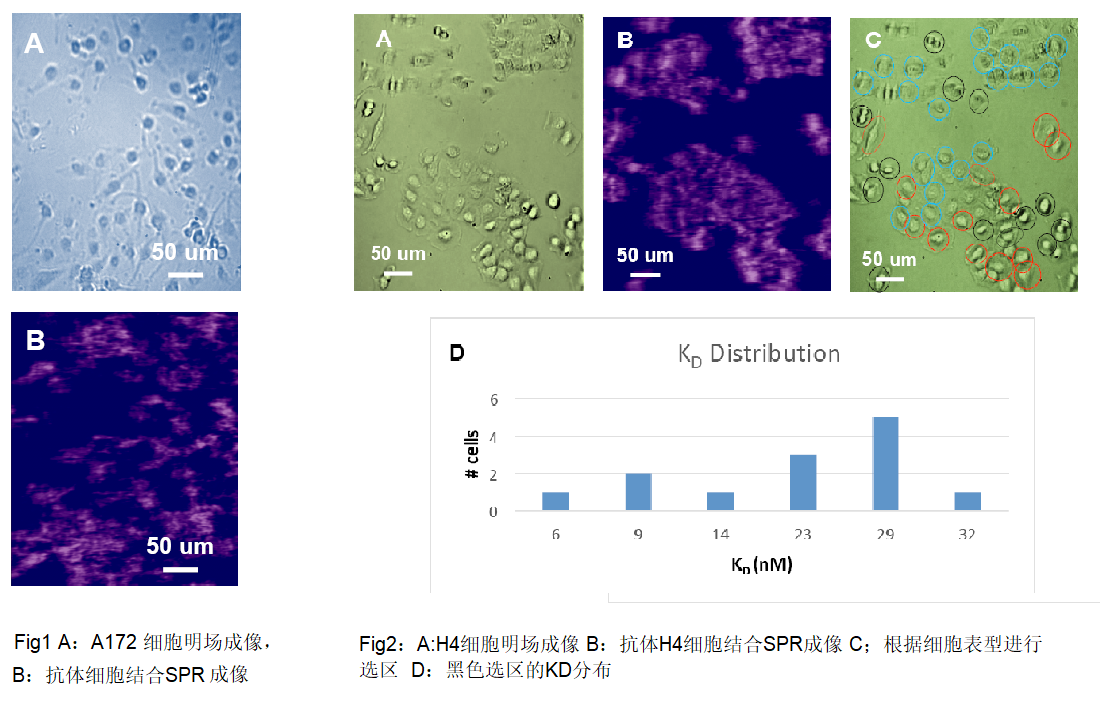
Study based on interaction between virus
and bacterial carrier molecules
1.Rapid ASTs experiment:study on
metabolic activity of antibiotics and escherichia coli(Live E.Coli O157.H7)

ASTs is an important
experiment to determine the antibiotic susceptibility and bacterial drug
resistance.Currently, most AST experiments are based on cell culture and take
several days to complete.Rapid AST detection can reduce morbidity and mortality
and help develop early narrow-spectrum antibiotic therapy. Through SPRm200, we
use plasma imaging and PIT technology to track the movement of individual
bacterial cells and monitor z-direction
changes with cell metabolism and antibiotic inputs. As can be seen, antibiotics
can significantly reduce the activity of bacterial cells and be calculated quantitatively, thus becoming a
fast AST detection method.
2.Study on
interaction between different GPCR receptors and ligands based on SPRm200 and
viral microarray

By electrochemical SPRM impedance analysis, the binding kinetic constants of viral peptide ligands and different GPCR receptors on the surface of the sensor were measured
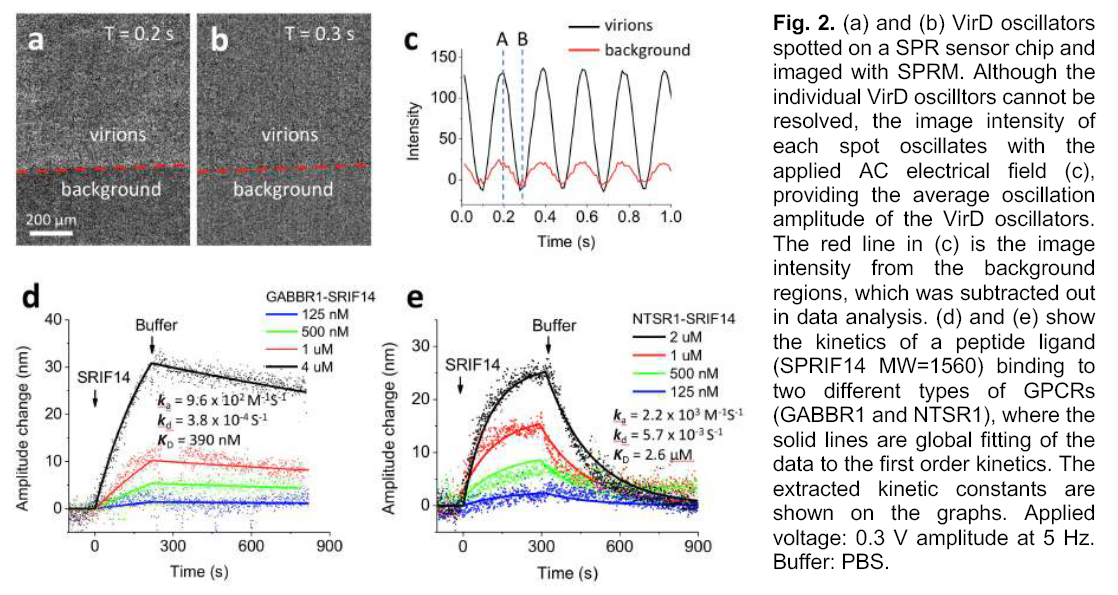
Papers(Part)
[1]
H Yu, X Shan, S Wang, N Tao, Achieving high spatial resolution surface plasmon
resonance microscopy with image reconstruction, Anal. Chem., 2017, 89 (5),
pp 2704–2707.
[2]
Y Wang, X Shan, H Wang, S Wang, N Tao, Plasmonic imaging of surface
electrochemical reactions of single gold nanowires, J. Am. Chem. Soc.,
2017, 139 (4), pp 1376–1379.
[3]
Dan Jiang, Yingyan Jiang, Zhimin Li, Tao Liu, Xiang Wo, Yimin Fang, Nongjian
Tao, Wei Wang, Hong-Yuan Chen, Optical imaging of phase transition and Li-ion
diffusion kinetics of single LiCoO2 nanoparticles during electrochemical
cycling, J. Am. Chem. Soc., 2017, 139 (1), pp 186–192.
[4]
Jin Lu, Yunze Yang, Wei Wang, Jinghong Li, Nongjian Tao, Shaopeng Wang,
Label-free imaging of histamine mediated G protein-coupled receptors activation
in live cells, Anal. Chem. 2016, 88, 11498−11503.
[5]
Karan Syal, Rafael Iriya, Yunze Yang, Hui Yu, Shaopeng Wang, Shelley E Haydel,
Hong-Yuan Chen, and Nongjian Tao, Antimicrobial Susceptibility Test with
Plasmonic Imaging and Tracking of Single Bacterial Motions on Nanometer Scale, ACS
Nano, 2016, 10(1), 845-852.
[6]
Fenni Zhang, Shaopeng Wang, Linliang Yin, Yunze Yang, Yan Guan, Wei Wang, Han
Xu and Nongjian Tao, Quantification of Epidermal Growth Factor Receptor
Expression Level and Binding Kinetics on Cell Surfaces by Surface Plasmon
Resonance Imaging, Anal. Chem. 2015, 87 (19), 9960-9965.
[7]
Linliang Yin, Yunze Yang, Shaopeng Wang, Wei Wang, Shengtao Zhang, Nongjian
Tao, Measuring the binding kinetics of antibody-conjugated gold nanoparticles
with intact cells, Small, 11(31), 3782-3788.
[8]
Yunze Yang, Hui Yu, Xiaonan Shan, Wei Wang, Xianwei Liu, Shaopeng
Wang, and Nongjian Tao, Label-Free Tracking of Single Organelle Transportation
in Cells with Nanometer Precision Using a Plasmonic Imaging Technique, Small, 11(24), 2878-2884, 2015
[8]
Linliang Yin, Wei Wang, Shaopeng Wang, Fenni Zhang, Shengtao Zhang, Nongjian
Tao, How does fluorescent labeling affect the binding kinetics of proteins with
intact cells? Biosensors and Bioelectronics, 2015, 66, 412-416.
[9]
Y Fang, S Chen, W Wang, X Shan, N Tao, Real-Time Monitoring of Phosphorylation
Kinetics with Self-Assembled Nano-oscillators, Angewandte Chemie
International Edition 2015, 54 (8), 2538-2542
[10]
Karan Syal, Wei Wang, Xiaonan Shan, Shaopeng Wang, Hong-Yuan Chen, Nongjian
Tao, Plasmonic imaging of protein interactions with single bacterial cells, Biosens.
Bioelectron., 2015, 63, 131-137.
[11]
Wei Wang, Linliang Yin, Laura Gonzalez-Malerva, Shaopeng Wang, Xiaobo Yu, Seron
Eaton, Shengtao Zhang, Hong-Yuan Chen, Joshua LaBaer, Nongjian Tao, In situ
drug-receptor binding kinetics in single cells: a quantitative label-free study
of anti-tumor drug resistance, Scientific Reports, 2014, 4
[12]
Hui Yu, Xiaonan Shan, Shaopeng Wang, Hong-Yuan Chen, Nongjian Tao, Molecular
scale origin of surface plasmon resonance biosensors, Analytical Chemistry,
2014, 86 (18), 8992-8997.
[13]
Xiaonan Shan, Yimin Fang, Shaopeng Wang, Yan Guan, Jongyuan Chen and Nongjian
Tao, Detection of charges and molecules with self-assembled nano-oscillators, Nano
Lett., 2014, 14 (7), 4151–4157.
[14]
Hui Yu, Xiaonan Shan, Shaopeng Wang, Jongyuan Chen and Nongjian Tao, Plasmonic
Imaging and Detection of Single DNA Molecules, ACS Nano, 2014, 8 (4),
3427–3433.
[15]
Xiaonan Shan, Ismael Díez-Pérez, Luojia Wang, Peter Wiktor, Ying Gu, Lihua
Zhang, Wei Wang, Jin Lu, Shaopeng Wang, Qihuang Gong, Jinghong Li &
Nongjian Tao, Imaging the electrocatalytic activity of single nanoparticles, Nature
Nanotechnology, 2012, 7, 668–672.
[16]
Wei Wang, Yunze Yang, Shaopeng Wang, Vinay J Nagaraj, Qiang Liu, Jie Wu and
Nongjian Tao, Label-free measuring and mapping of binding kinetics of membrane
proteins in single living cells, Nature Chemistry, 2012, 4(10),
846-53.
[17]
Wang, Wei; Wang, Shaopeng; Liu, Qiang; Wu, Jie; Tao, Nongjian, Mapping
Single-Cell–Substrate Interactions by Surface Plasmon Resonance Microscopy, Langmuir,
2012, 28(37), 13373-13379.
[18]
S. Wang, X. Shan, U. Patel, X. Huang, J. Lu, J. Li, NJ Tao, Label-free imaging,
detection and mass measurement of single viruses by Surface Plasmon Resonance, Proc
Natl Acad Sci U S A, 2010, 107 (37), 16028-16032













